242
Members
11
Research Team
75
Researchers
69
Engineers, Technicians, Administrative staff
CIIL'S RECENT PUBLICATIONS
-
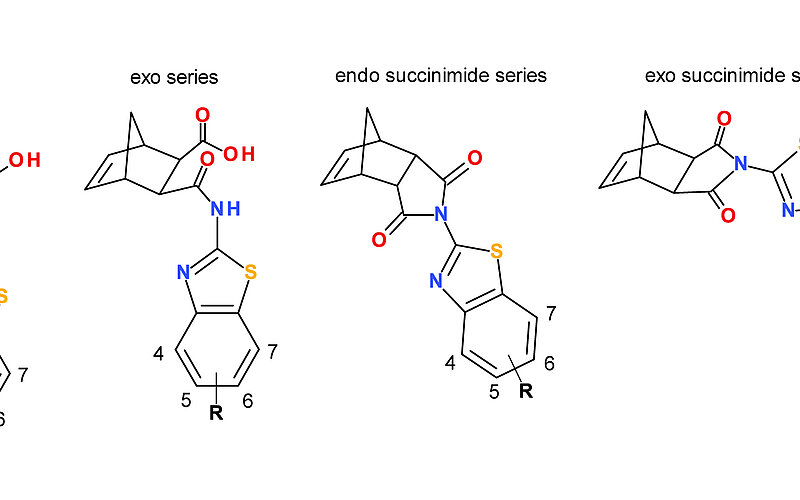
Exploiting DprE2 pathway redundancy to enhance potency and overcome resistance to pretomanid in…
-

SphB1 export system for antigen presentation to the respiratory tract by a live pertussis…
-
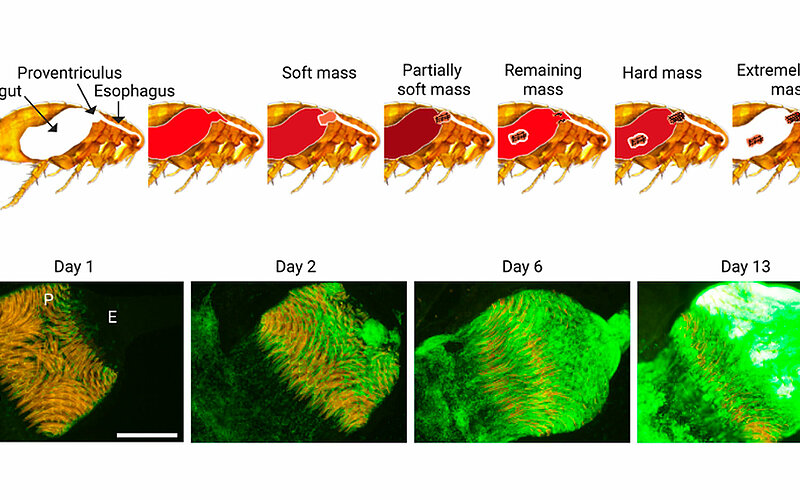
Metabolic adaptation of Yersinia pestis during flea colonization
-
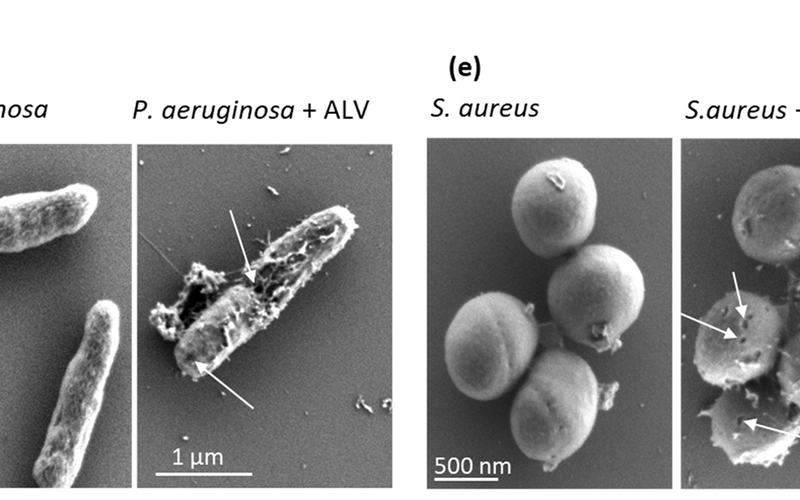

An Extremophile Deep Sea Peptide to Combat Cystic Fibrosis Infections Caused by Pseudomonas…
-
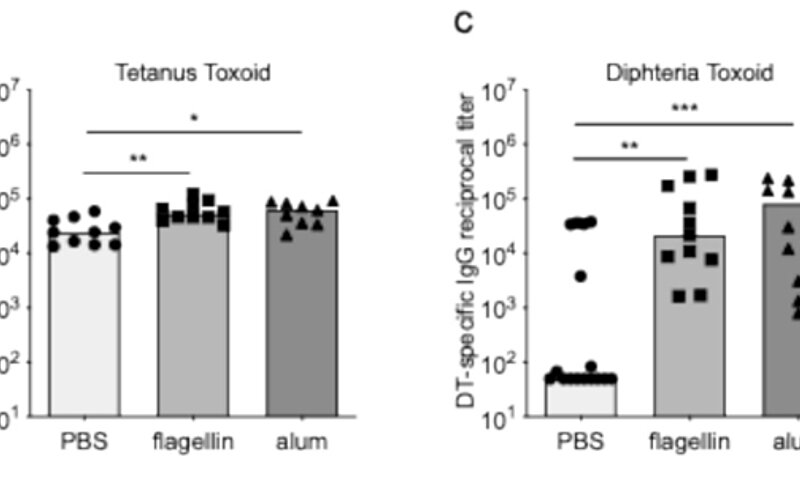
Toll-like receptor 5 agonist is a potent adjuvant for intradermal vaccination.
CIIL'S LATEST NEWS
-
 Francophone Congress of Allergology 2026, at the Palais des Congrès de Paris
Francophone Congress of Allergology 2026, at the Palais des Congrès de Paris -
 An interview with Dr. Inès Leleu, a postdoctoral researcher at the Lille Center for Infection and Immunity and a member of the ERA4TB project.
An interview with Dr. Inès Leleu, a postdoctoral researcher at the Lille Center for Infection and Immunity and a member of the ERA4TB project. -
 Ideas, talent and community: our 2025 off-site day
Ideas, talent and community: our 2025 off-site day -
 Engaged in field research with Dr. Fernando Real
Engaged in field research with Dr. Fernando Real -
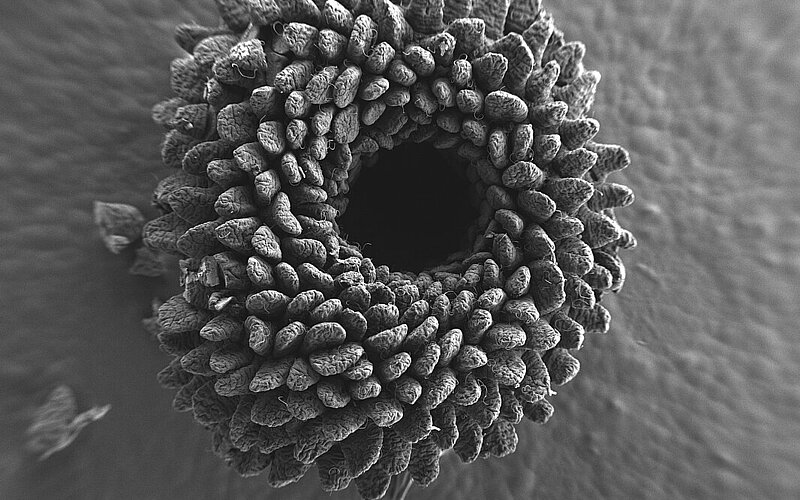 Nicolas Barois, affiliated with CIIL U1019 (UMR 9017) / BICeL Platform, UAR 2014 – US 41, PLBS (Lille Platforms in Biology and Health), has been awarded First Prize in the 2025 FBI Image Contest.
Nicolas Barois, affiliated with CIIL U1019 (UMR 9017) / BICeL Platform, UAR 2014 – US 41, PLBS (Lille Platforms in Biology and Health), has been awarded First Prize in the 2025 FBI Image Contest. -
 The awarding of the poster winner has been attributed to Joan while the 7th European Congress of Immunology
The awarding of the poster winner has been attributed to Joan while the 7th European Congress of Immunology -
 Success stories
Success stories












